One-to-n alignments
Command: compare-matrices -v 1 -mode matches -format1 transfac -file1 $RSAT/public_html/tmp/wwwrun/2014/09/03/peak-motifs.2014-09-03.182940_2014-09-03.182940_LkVj4M/results/discovered_motifs/oligos_6nt_mkv4_m5/peak-motifs_oligos_6nt_mkv4_m5.tf -format2 tf -file2 $RSAT/public_html/tmp/wwwrun/2014/09/03/peak-motifs.2014-09-03.182940_2014-09-03.182940_LkVj4M/peak-motifs_custom_motif_db.tf -mode matches -DR -uth offset_rank 1 -lth w 5 -lth Wr 0.3 -lth cor 0.75 -lth Ncor 0.4 -return matrix_name,matrix_id,cor,Ncor,logoDP,NIcor,NsEucl,SSD,NSW,match_rank,width,strand,offset,consensus,alignments_1ton -sort Ncor -o $RSAT/public_html/tmp/wwwrun/2014/09/03/peak-motifs.2014-09-03.182940_2014-09-03.182940_LkVj4M/results/discovered_motifs/oligos_6nt_mkv4_m5/peak-motifs_oligos_6nt_mkv4_m5_vs_db_Schmidt_CEBPa_motif
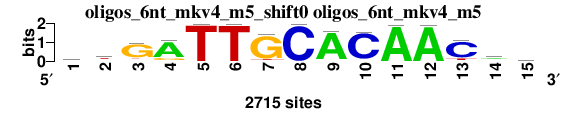
One-to-n matrix alignment; reference matrix: oligos_6nt_mkv4_m5_shift0 ; 2 matrices ; sort_field=rank_mean
| Matrix name | Aligned logos | cor |
Ncor |
logoDP |
NIcor |
NsEucl |
SSD |
NSW |
rcor |
rNcor |
rlogoDP |
rNIcor |
rNsEucl |
rSSD |
rNSW |
rank_mean |
match_rank |
Aligned matrices |
|---|
| oligos_6nt_mkv4_m5_shift0 (oligos_6nt_mkv4_m5) |
 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
; oligos_6nt_mkv4_m5; m=0 (reference); ncol1=15; shift=0; ncol=15; atGATTGCACAACwy
; Alignment reference
a 1046 663 145 2141 1 1 72 6 2545 94 2687 2687 42 1152 677
c 471 240 158 56 4 1 6 2698 51 2522 26 10 2208 613 803
g 578 483 2160 474 4 10 2428 2 31 16 1 9 95 166 539
t 620 1329 252 44 2706 2703 209 9 88 83 1 9 370 784 696
|
| do560+do843_mmus_cebpa_liver_meme_top_m1_shift3 (do560+do843_mmus_cebpa_liver_meme_top_m1) |
 |
0.926 |
0.679 |
9.366 |
0.663 |
0.937 |
0.962 |
0.956 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1.000 |
1 |
; oligos_6nt_mkv4_m5 versus do560+do843_mmus_cebpa_liver_meme_top_m1; m=1/1; ncol2=11; w=11; offset=3; strand=D; shift=3; score= 1; ---rTTGCacMAym-
; cor=0.926; Ncor=0.679; logoDP=9.366; NIcor=0.663; NsEucl=0.937; SSD=0.962; NSW=0.956; rcor=1; rNcor=1; rlogoDP=1; rNIcor=1; rNsEucl=1; rSSD=1; rNSW=1; rank_mean=1.000; match_rank=1
a 0 0 0 204 0 0 35 0 259 80 300 431 1 143 0
c 0 0 0 69 0 0 0 431 48 258 130 0 169 145 0
g 0 0 0 138 0 0 365 0 85 0 0 0 78 60 0
t 0 0 0 20 431 431 31 0 39 93 1 0 183 83 0
|

