One-to-n alignments
Command: compare-matrices -v 1 -mode matches -format1 transfac -file1 $RSAT/public_html/tmp/apache/2014/09/05/peak-motifs.2014-09-05.194903_2014-09-05.194903_oebJl9/results/discovered_motifs/positions_8nt_m3/peak-motifs_positions_8nt_m3.tf -format2 tf -file2 $RSAT/public_html/data/motif_databases/JASPAR/jaspar_core_vertebrates_2013-11.tf -mode matches -DR -uth offset_rank 1 -lth w 5 -lth Wr 0.3 -lth cor 0.75 -lth Ncor 0.4 -return matrix_name,matrix_id,cor,Ncor,logoDP,NIcor,NsEucl,SSD,NSW,match_rank,width,strand,offset,consensus,alignments_1ton -sort Ncor -o $RSAT/public_html/tmp/apache/2014/09/05/peak-motifs.2014-09-05.194903_2014-09-05.194903_oebJl9/results/discovered_motifs/positions_8nt_m3/peak-motifs_positions_8nt_m3_vs_db_jaspar_core_vertebrates
One-to-n matrix alignment; reference matrix: positions_8nt_m3_shift0 ; 3 matrices ; sort_field=rank_mean
| Matrix name | Aligned logos | cor |
Ncor |
logoDP |
NIcor |
NsEucl |
SSD |
NSW |
rcor |
rNcor |
rlogoDP |
rNIcor |
rNsEucl |
rSSD |
rNSW |
rank_mean |
match_rank |
Aligned matrices |
|---|
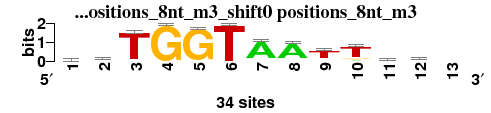
| positions_8nt_m3_shift0 (positions_8nt_m3) |
 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
; positions_8nt_m3; m=0 (reference); ncol1=12; shift=0; ncol=13; dsTGGTAATTcw-
; Alignment reference
a 10 6 1 0 0 0 29 28 5 3 7 13 0
c 3 13 0 0 0 0 2 2 2 1 15 8 0
g 10 12 1 34 33 0 2 2 3 4 7 2 0
t 11 3 32 0 1 34 1 2 24 26 5 11 0
|
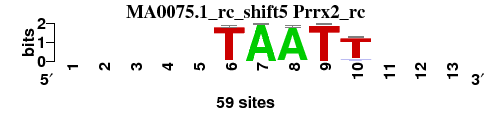
| MA0075.1_rc_shift5 (Prrx2_rc) |
 |
0.983 |
0.410 |
1.577 |
0.069 |
0.935 |
0.211 |
0.979 |
1 |
2 |
2 |
2 |
1 |
1 |
1 |
1.429 |
1 |
; positions_8nt_m3 versus MA0075.1_rc (Prrx2_rc); m=1/2; ncol2=5; w=5; offset=5; strand=R; shift=5; score= 1.4286; -----TAATT---
; cor=0.983; Ncor=0.410; logoDP=1.577; NIcor=0.069; NsEucl=0.935; SSD=0.211; NSW=0.979; rcor=1; rNcor=2; rlogoDP=2; rNIcor=2; rNsEucl=1; rSSD=1; rNSW=1; rank_mean=1.429; match_rank=1
a 0 0 0 0 0 0 59 58 0 1 0 0 0
c 0 0 0 0 0 1 0 1 0 4 0 0 0
g 0 0 0 0 0 0 0 0 0 2 0 0 0
t 0 0 0 0 0 58 0 0 59 52 0 0 0
|
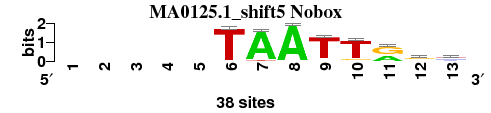
| MA0125.1_shift5 (Nobox) |
 |
0.877 |
0.472 |
3.962 |
0.501 |
0.909 |
0.807 |
0.942 |
2 |
1 |
1 |
1 |
2 |
2 |
2 |
1.571 |
2 |
; positions_8nt_m3 versus MA0125.1 (Nobox); m=2/2; ncol2=8; w=7; offset=5; strand=D; shift=5; score= 1.5714; -----TAATTrsy
; cor=0.877; Ncor=0.472; logoDP=3.962; NIcor=0.501; NsEucl=0.909; SSD=0.807; NSW=0.942; rcor=2; rNcor=1; rlogoDP=1; rNIcor=1; rNsEucl=2; rSSD=2; rNSW=2; rank_mean=1.571; match_rank=2
a 0 0 0 0 0 0 36 38 1 2 15 4 2
c 0 0 0 0 0 1 0 0 1 0 0 12 13
g 0 0 0 0 0 0 0 0 2 4 22 18 6
t 0 0 0 0 0 37 2 0 34 32 1 4 17
|


