One-to-n alignments
Command: compare-matrices -v 1 -mode matches -format1 transfac -file1 $RSAT/public_html/tmp/apache/2014/09/05/peak-motifs.2014-09-05.202657_2014-09-05.202657_8OTbPR/results/discovered_motifs/dyads_m2/peak-motifs_dyads_m2.tf -format2 tf -file2 $RSAT/public_html/data/motif_databases/footprintDB/footprintDB.motif.tf -mode matches -DR -uth offset_rank 1 -lth w 5 -lth Wr 0.3 -lth cor 0.75 -lth Ncor 0.4 -return matrix_name,matrix_id,cor,Ncor,logoDP,NIcor,NsEucl,SSD,NSW,match_rank,width,strand,offset,consensus,alignments_1ton -sort Ncor -o $RSAT/public_html/tmp/apache/2014/09/05/peak-motifs.2014-09-05.202657_2014-09-05.202657_8OTbPR/results/discovered_motifs/dyads_m2/peak-motifs_dyads_m2_vs_db_footprintDB
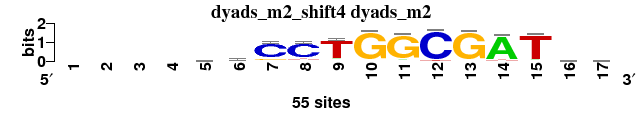
One-to-n matrix alignment; reference matrix: dyads_m2_shift4 ; 4 matrices ; sort_field=rank_mean
| Matrix name | Aligned logos | cor |
Ncor |
logoDP |
NIcor |
NsEucl |
SSD |
NSW |
rcor |
rNcor |
rlogoDP |
rNIcor |
rNsEucl |
rSSD |
rNSW |
rank_mean |
match_rank |
Aligned matrices |
|---|
| dyads_m2_shift4 (dyads_m2) |
 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
; dyads_m2; m=0 (reference); ncol1=13; shift=4; ncol=17; ----mkCCTGGCGATwm
; Alignment reference
a 0 0 0 0 17 7 1 3 4 0 3 0 2 50 3 15 15
c 0 0 0 0 15 11 44 45 1 1 0 52 0 1 2 13 15
g 0 0 0 0 10 16 7 3 3 52 50 0 52 1 0 10 12
t 0 0 0 0 13 21 3 4 47 2 2 3 1 3 50 17 13
|
| 4955_brk_DrosophilaTF_1.1__shift8 (4955_brk_DrosophilaTF_1.1_) |
 |
0.835 |
0.449 |
7.137 |
0.470 |
0.881 |
1.389 |
0.901 |
2 |
1 |
1 |
1 |
2 |
2 |
2 |
1.571 |
1 |
; dyads_m2 versus 4955_brk_DrosophilaTF_1.1_; m=1/3; ncol2=7; w=7; offset=4; strand=D; shift=8; score= 1.5714; --------TGGCGyY--
; cor=0.835; Ncor=0.449; logoDP=7.137; NIcor=0.470; NsEucl=0.881; SSD=1.389; NSW=0.901; rcor=2; rNcor=1; rlogoDP=1; rNIcor=1; rNsEucl=2; rSSD=2; rNSW=2; rank_mean=1.571; match_rank=1
a 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0 0
c 0 0 0 0 0 0 0 0 0 0 0 1 0 0.5833 0.25 0 0
g 0 0 0 0 0 0 0 0 0.0833 1 1 0 1 0 0 0 0
t 0 0 0 0 0 0 0 0 0.9167 0 0 0 0 0.4167 0.75 0 0
|
| 2843_PF0156.1_JASPAR_CORE_2009__rc_shift0 (2843_PF0156.1_JASPAR_CORE_2009__rc) |
 |
0.839 |
0.444 |
0.356 |
-0.083 |
0.908 |
1.382 |
0.923 |
1 |
2 |
3 |
3 |
1 |
1 |
1 |
1.714 |
2 |
; dyads_m2 versus 2843_PF0156.1_JASPAR_CORE_2009__rc; m=2/3; ncol2=13; w=9; offset=-4; strand=R; shift=0; score= 1.7143; GCGsCmcCTGGyG----
; cor=0.839; Ncor=0.444; logoDP=0.356; NIcor=-0.083; NsEucl=0.908; SSD=1.382; NSW=0.923; rcor=1; rNcor=2; rlogoDP=3; rNIcor=3; rNsEucl=1; rSSD=1; rNSW=1; rank_mean=1.714; match_rank=2
a 0 0 0 17 0 103 16 49 0 0 0 0 39 0 0 0 0
c 0 292 0 152 292 138 169 243 0 0 0 175 0 0 0 0 0
g 292 0 292 95 0 47 68 0 0 292 292 0 253 0 0 0 0
t 0 0 0 28 0 4 39 0 292 0 0 117 0 0 0 0 0
|
| 2606_MA0368.1_JASPAR_CORE_2009__rc_shift6 (2606_MA0368.1_JASPAR_CORE_2009__rc) |
 |
0.760 |
0.409 |
1.735 |
-0.030 |
0.855 |
2.052 |
0.853 |
3 |
3 |
2 |
2 |
3 |
3 |
3 |
2.714 |
3 |
; dyads_m2 versus 2606_MA0368.1_JASPAR_CORE_2009__rc; m=3/3; ncol2=7; w=7; offset=2; strand=R; shift=6; score= 2.7143; ------CTTGGCr----
; cor=0.760; Ncor=0.409; logoDP=1.735; NIcor=-0.030; NsEucl=0.855; SSD=2.052; NSW=0.853; rcor=3; rNcor=3; rlogoDP=2; rNIcor=2; rNsEucl=3; rSSD=3; rNSW=3; rank_mean=2.714; match_rank=3
a 0 0 0 0 0 0 0 0 0 0 0 0 39 0 0 0 0
c 0 0 0 0 0 0 100 0 0 0 0 100 9 0 0 0 0
g 0 0 0 0 0 0 0 0 0 100 100 0 43 0 0 0 0
t 0 0 0 0 0 0 0 100 100 0 0 0 9 0 0 0 0
|



